Proteomic Identification of protease Cleavage Sites (PICS) workflow.... | Download Scientific Diagram

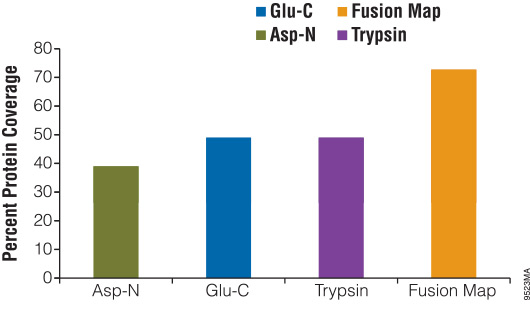
Peptide mapping of MPI molecule using Glu-C protease. A 50 l g sample... | Download Scientific Diagram
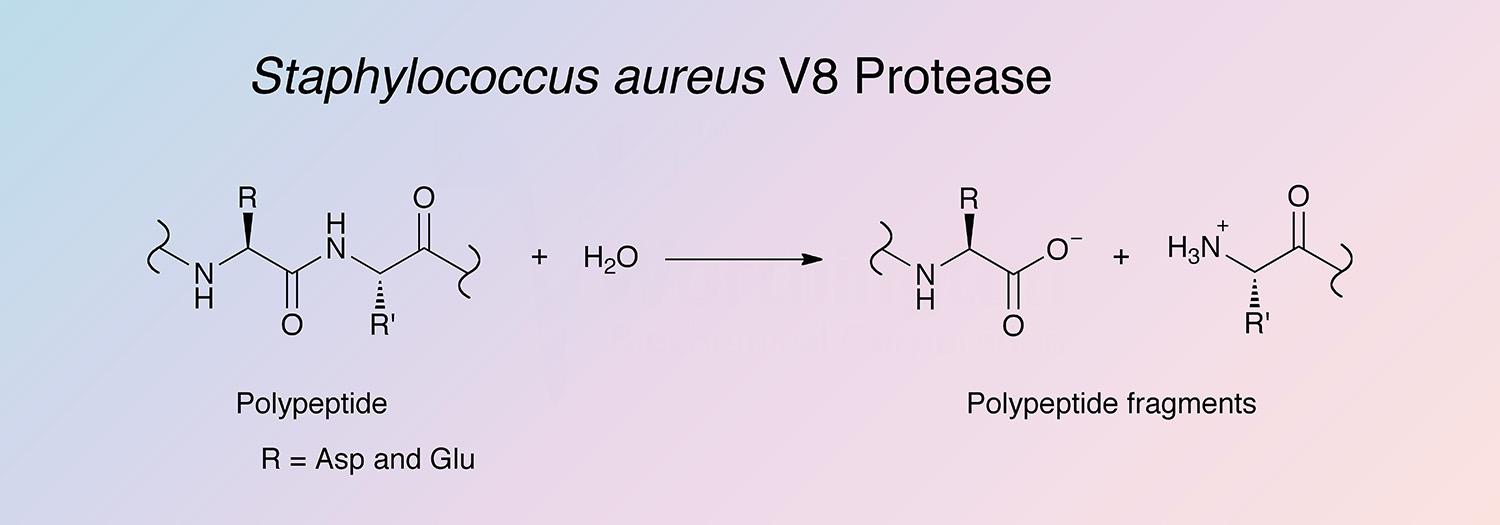
Buy Recombinant V8 Protease (Glu-C) Pharmacy Grade Pharmacy Grade from Shanghai Yaxin Biotechnology Co., Ltd. - ECHEMI

MP Biomedicals™ Protease (S. Aureus) V.8 5mg Proteases and Protein-Cleaving Reagents | Fisher Scientific
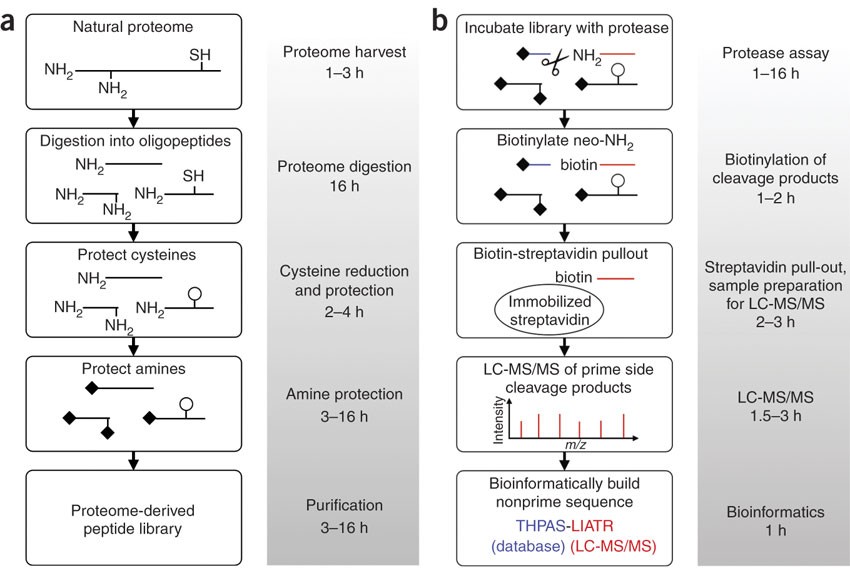
Proteome-derived, database-searchable peptide libraries for identifying protease cleavage sites | Nature Biotechnology
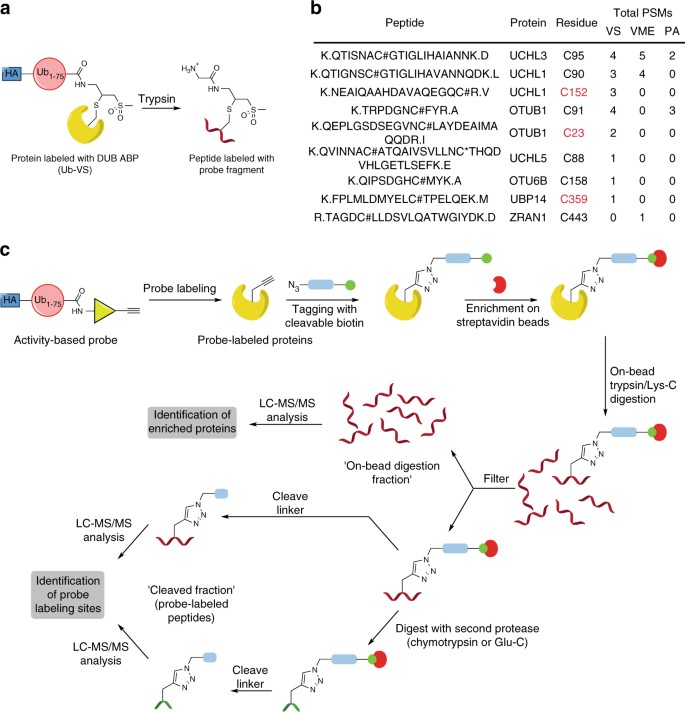
Reactive-site-centric chemoproteomics identifies a distinct class of deubiquitinase enzymes | Nature Communications